- Cytochrome c oxidase
-
Cytochrome-c oxidase 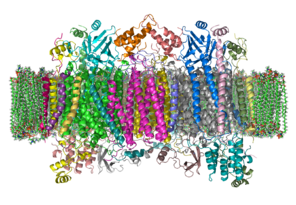
The crystal structure of bovine cytochrome c oxidase in a phospholipid bilayer. The intermembrane space lies to top of the image. Adapted from PDB 1OCC (It is a homo dimer in this structure) Identifiers EC number 1.9.3.1 CAS number 9001-16-5 Databases IntEnz IntEnz view BRENDA BRENDA entry ExPASy NiceZyme view KEGG KEGG entry MetaCyc metabolic pathway PRIAM profile PDB structures RCSB PDB PDBe PDBsum Gene Ontology AmiGO / EGO Search PMC articles PubMed articles The enzyme cytochrome c oxidase or Complex IV (PDB 2OCC, EC 1.9.3.1) is a large transmembrane protein complex found in bacteria and the mitochondrion.
It is the last enzyme in the respiratory electron transport chain of mitochondria (or bacteria) located in the mitochondrial (or bacterial) membrane. It receives an electron from each of four cytochrome c molecules, and transfers them to one oxygen molecule, converting molecular oxygen to two molecules of water. In the process, it binds four protons from the inner aqueous phase to make water, and in addition translocates four protons across the membrane, helping to establish a transmembrane difference of proton electrochemical potential that the ATP synthase then uses to synthesize ATP.
Contents
Structure
The complex is a large integral membrane protein composed of several metal prosthetic sites and 13 protein subunits in mammals. In mammals, ten subunits are nuclear in origin, and three are synthesized in the mitochondria. The complex contains two hemes, a cytochrome a and cytochrome a3, and two copper centers, the CuA and CuB centers.[1] In fact, the cytochrome a3 and CuB form a binuclear center that is the site of oxygen reduction. Cytochrome c reduced by the preceding component of the respiratory chain (cytochrome bc1 complex, complex III) docks near the CuA binuclear center, passing an electron to it and being oxidized back to cytochrome c containing Fe3+. The reduced CuA binuclear center now passes an electron on to cytochrome a, which in turn passes an electron on to the cytochrome a3- CuB binuclear center. The two metal ions in this binuclear center are 4.5 Å apart and coordinate a hydroxide ion in the fully oxidized state.
Crystallographic studies of cytochrome c oxidase show an unusual post-translational modification, linking C6 of Tyr(244) and the ε-N of His(240) (bovine enzyme numbering). It plays a vital role in enabling the cytochrome a3- CuB binuclear center to accept four electrons in reducing molecular oxygen to water. The mechanism of reduction was formerly thought to involve a peroxide intermediate, which was believed to lead to superoxide production. However, the currently accepted mechanism involves a rapid four electron reduction involving immediate oxygen-oxygen bond cleavage, avoiding any intermediate likely to form superoxide.[2]
Assembly
Site of assembly is believed to occur near TOM/TIM, where complex intermediates are accessible to bind with subunits imported from cytosol. Hemes and cofactors are inserted into subunits I & II Subunits I and IV initiate assembly. Other subunits may form sub-complex intermediates that later bind to others to form COX complex. In post-assembly modifications, the enzyme is dimerized, which is required for active/efficient enzyme action. Dimers are connected by a cardiolipin molecule.[3][4]
Biochemistry
Summary reaction:
- 4 Fe2+-cytochrome c + 8 H+in + O2 → 4 Fe3+-cytochrome c + 2 H2O + 4 H+out
Two electrons are passed from two cytochrome c's, through the CuA and cytochrome a sites to the cytochrome a3- CuB binuclear center, reducing the metals to the Fe+2 form and Cu+1. The hydroxide ligand is protonated and lost as water, creating a void between the metals which is filled by O2. The oxygen is rapidly reduced, with two electrons coming from the Fe+2cytochrome a3, which is converted to the ferryl oxo form (Fe+4=O). The oxygen atom close to CuB picks up one electron from Cu+1, and a second electron and a proton from the hydroxyl of Tyr(244), which becomes a tyrosyl radical: the second oxygen is converted to a hydroxide ion by picking up two electrons and a proton. A third electron arising from another cytochrome c is passed through the first two electron carriers to the cytochrome a3- CuB binuclear center, and this electron and two protons convert the tyrosyl radical back to Tyr, and the hydroxide bound to CuB+2 to a water molecule. The fourth electron from another cytochrome c flows through CuA and cytochrome a to the cytochrome a3- CuB binuclear center, reducing the Fe+4=O to Fe+3, with the oxygen atom picking up a proton simultaneously, regenerating this oxygen as a hydroxide ion coordinated in the middle of the cytochrome a3- CuB center as it was at the start of this cycle. The net process is that four reduced cytochrome c's are used, along with 4 protons, to reduce O2 to two water molecules.
Inhibition
Cyanide, sulfide, azide, and carbon monoxide[5] all bind to cytochrome c oxidase, thus competitively inhibiting the protein from functioning which results in chemical asphyxiation of cells. Methanol in methylated spirits is converted into formic acid which also inhibits the same oxidase system.
Genetic defects and disorders
Defects involving genetic mutations altering cytochrome c oxidase (COX) functionality or structure can result in severe, often fatal metabolic disorders. Such disorders usually manifest in early childhood and predominantly affect tissues with high energy demands (brain, heart, muscle). Among the many classified mitochondrial diseases, those involving dysfunctional COX assembly are thought to be the most severe.[6]
The vast majority of COX disorders are linked to mutations in nuclear-encoded proteins referred to as assembly factors, or assembly proteins. These assembly factors contribute to COX structure and functionality, and are involved in several essential processes, including transcription and translation of mitochondrion-encoded subunits, processing of preproteins and membrane insertion, and cofactor biosynthesis and incorporation.[7]
Currently, mutations have been identified in six COX assembly factors: SURF1, SCO1, SCO2, COX10, COX15, and LRPPRC. Mutations in these proteins can result in altered functionality of sub-complex assembly, copper transport, or translational regulation. Each gene mutation is associated with the etiology of a specific disease, with some having implications in multiple disorders. Disorders involving dysfunctional COX assembly via gene mutations include Leigh syndrome, cardiomyopathy, leukodystrophy, anemia, and sensorineural deafness.
Histochemistry
COX histochemistry is used for mapping regional brain metabolism in animals, since there is a direct relation between enzyme activity and neuronal activity.[8] Such brain mapping has been accomplished in spontaneous mutant mice with cerebellar disease such as reeler[9] and a transgenic model of Alzheimer's disease.[10] This technique has also been used to map learning activity in animal brain.[11]
Additional images
See also
- Cytochrome c oxidase subunit I
- Cytochrome c oxidase subunit II
- Cytochrome c oxidase subunit III
- Heme a
References
- ^ Tsukihara T, Aoyama H, Yamashita E, Tomizaki T, Yamaguchi H, Shinzawa-Itoh K, Nakashima R, Yaono R, Yoshikawa S (August 1995). "Structures of metal sites of oxidized bovine heart cytochrome c oxidase at 2.8 A". Science 269 (5227): 1069–74. doi:10.1126/science.7652554. PMID 7652554.
- ^ Voet, Donald (2010). Biochemistry. New York: J. Wiley & Sons. pp. 865–866. ISBN 0-470-57095-4.
- ^ Khalimonchuk O, Rödel G (December 2005). "Biogenesis of cytochrome c oxidase". Mitochondrion 5 (6): 363–88. doi:10.1016/j.mito.2005.08.002. PMID 16199211.
- ^ Fontanesi F, Soto IC, Horn D, Barrientos A (December 2006). "Assembly of mitochondrial cytochrome c-oxidase, a complicated and highly regulated cellular process". Am. J. Physiol., Cell Physiol. 291 (6): C1129–47. doi:10.1152/ajpcell.00233.2006. PMID 16760263.
- ^ Alonso JR, Cardellach F, López S, Casademont J, Miró O (September 2003). "Carbon monoxide specifically inhibits cytochrome c oxidase of human mitochondrial respiratory chain". Pharmacol. Toxicol. 93 (3): 142–6. doi:10.1034/j.1600-0773.2003.930306.x. PMID 12969439.
- ^ Pecina P, Houstková H, Hansíková H, Zeman J, Houstek J (2004). "Genetic defects of cytochrome c oxidase assembly". Physiol Res 53 Suppl 1: S213–23. PMID 15119951. http://www.biomed.cas.cz/physiolres/pdf/53%20Suppl%201/53_S213.pdf.
- ^ Zee JM, Glerum DM (December 2006). "Defects in cytochrome oxidase assembly in humans: lessons from yeast". Biochem. Cell Biol. 84 (6): 859–69. doi:10.1139/o06-201. PMID 17215873.
- ^ Wong-Riley MT. (1989). "Cytochrome oxidase: an endogenous metabolic marker for neuronal activity.". Trends Neurosci. 12 (3): 94-111. PMID 2469224.
- ^ Strazielle C, Hayzoun K, Derer M, Mariani J, Lalonde R. (April 2006). "Regional brain variations of cytochrome oxidase activity in Relnrl-orl mutant mice.". J. Neurosci. Res. 83 (5): 821–31. PMID 16511878.
- ^ Strazielle C, Sturchler-Pierrat C, Staufenbiel M, Lalonde R. (2003). "Regional brain cytochrome oxidase activity in beta-amyloid precursor protein transgenic mice with the Swedish mutation.". Neuroscience 118 (4): 1151-63. PMID 12732258.
- ^ Conejo NM, González-Pardo H, Gonzalez-Lima F, Arias JL. (2010). "Spatial learning of the water maze: progression of brain circuits mapped with cytochrome oxidase histochemistry.". Neurobiol. Learn. Mem. 93 (3): 362-71. PMID 19969098.
External links
- The Cytochrome Oxidase home page at Rice University
- Interactive Molecular model of cytochrome c oxidase (Requires MDL Chime)
- UMich Orientation of Proteins in Membranes families/superfamily-4
- MeSH Cytochrome-c+Oxidase
Metabolism: Citric acid cycle enzymes Cycle Anaplerotic to acetyl-CoAPyruvate dehydrogenase complex (E1, E2, E3)
(regulated by Pyruvate dehydrogenase kinase and Pyruvate dehydrogenase phosphatase)to succinyl-CoAto oxaloacetateMitochondrial
electron transport chain/
oxidative phosphorylationPrimaryComplex I/NADH dehydrogenase · Complex II/Succinate dehydrogenase · Coenzyme Q · Complex III/Coenzyme Q - cytochrome c reductase · Cytochrome c · Complex IV/Cytochrome c oxidase
Coenzyme Q10 synthesis: COQ2 · COQ3 · COQ4 · COQ5 · COQ6 · COQ7 · COQ9 · COQ10A · COQ10B · PDSS1 · PDSS2OtherETC ETC Complex I - ETC Complex III - ETC Complex IVOther B memb: cead, trns (1A, 1C, 1F, 2A, 3A1, 3A2-3, 3D), othr Categories:- Cellular respiration
- EC 1.9.3
- Hemoproteins
- Integral membrane proteins
Wikimedia Foundation. 2010.




